Companies transitioning to or adopting Neo4j inevitably face the challenge of getting large amounts of (legacy) data into their new (knowledge) graph. This often entails a large discussion about how to organize things (aka the ontology or graph schema) and how to technically make it happen. The ontological aspect is on its own quite an important topic but this article focuses on the technical effort.
When ingesting large amounts of data (tens of Gb) into Neo4j there really is only one option: the neo4j-admin import utility. Everything else, including the ‘Load CSV‘ Cypher path is too slow to consider. The CLI import utility does the job but it also erases all existing data and you need a few post-fixes (restarting the database e.g.). Once the baseline is set the incremental data changes are usually discussed in the context of a CDC solution (change data capture) and this is where the non-destructive (Cypher-Bolt based) ingestion options come in. Companies often use cloud solutions (e.g. AWS Kinesis) or Apache Kafka for streaming data changes while static or almost static data changes require some form of scheduler and workflow platforms. This is where I advise customers to consider Apache Airflow, Apache Hop or messaging systems like RabbitMQ and alike.
Apache Airflow is a Python platform and this is also one of the main selling points for companies with a focus on data science (and/or a strong Python stack). Something like Apache Hop is an alternative but the typical Java context is often more difficult to digest for Python developers. Many customers like AWS Glue and other data platforms available on AWS or Azure, but the main disadvantage is the fact that these ETL platforms focus on marshaling relational or tabular data. Neo4j or similar graph backends are not very much supported by AWS Glue. So, when it comes to Neo4j ETL on premise or in the cloud (or both), Airflow is an ideal solution and one I like to advertise when companies request an ETL, CDC or inception solution.
The remains of this article describes the typical customer scenario:
- how to get tons of data into a brand new Neo4j database
- how to update the graph on regular intervals or explicitly when necessary
- how to approach Airflow development to create a robust ETL solution.
The focus here is on how to ingest data in Neo4j but the blueprint below really works with any data source and any endpoint. Airflow is capable of marshaling between lots of things, it’s very much a plumbing toolbox.
Before diving into the technicalities, a few words and thoughts on the good and the lesser of Airflow. Like all of the Apache tools and frameworks, Airflow has its quirks and open source challenges. It’s not polished and it comes with a learning curve. Many Apache frameworks (say Apache Hop or Apache Hadoop) are Java based and the fact that Airflow is fully Python helps a lot in understanding things, since one can peek at the implementation. The terminology is not very standard and other things can lead one astray. For example, a task is not an element of a flow but rather the flow itself while the term ‘operator’ is used for flow steps. The term DAG (direct acyclic graph) is used to refer to a flow. I think it would help a lot if Airflow would rephrase a few things.
On a UI level things are also rather mediocre, but that’s almost a trademark of Apache. The thing that remains rather incomprehensible is why one can’t upload/update a flow via the UI. Meaning that in order to manage/add flows one needs access to the underlying OS (directories). This, in a way, indicates how to develop things with/for Airflow: you need to develop things locally and hand it over to some admin once it’s ready. Airflow can’t really be used as a service backend. If there is a DAG set up it can be managed and the UI is effective but it fails miserably towards development purposes.
On the upside, Airflow can be used for anything and everything you wish to schedule. It can connect to anything and if you can program it in Python you can run it in Airflow. This includes things like reaching out to AWS SageMaker, triggering workflows based on directory changes (so-called sensors), ingesting any type of data and so on.
Airflow is NOT designed for streaming data, that’s where Apache Kafka comes in. Airflow data needs to be static. You can schedule things as often as you like but Airflow does not run hot.
Airflow does not replace some due diligence. To be specific, Airflow will run flows which can last days but it’s not a solution for poorly written code and poor performance. You need to research the various servers and services before trying to connect them. With respect to Neo4j, the classic Cypher loading approach takes days if you have tens of Gb but takes minutes via the import utility.
The main development challenge when creating an Airflow DAG is how to run and debug it:
- the essential thing to understand is that the flows in Airflow are scripts in a directory. By giving multiple people access to this you easily end up with clashing pip dependencies and custom functions.
- if you execute a DAG via the UI you will see log output but the cycle of copy-pasting new code in a directory, running the DAG and looking at the log is obviously not productive. It also means you have access to the DAG directory of Airflow.
- Airflow does not enforce a particular code organization and without it the management of flows can easily get out of hand
Towards a production-level ETL platform one needs:
- DAG templates, in order to have a uniform directory strucuture which can be managed and understood
- local development and unit tests prior to Airflow deployment
- CI/CD pipelines from Github (or alike) which take over the DAG, set up the (environment) variables, pip dependencies, connections and so on.
The inclusion of contextual elements (vars, connections….) is something the Astronomer solution does well and is unfortunately not part of Airflow. The crux here is that every flow depends on some settings which can (and should) be defined on the Airflow level (see below). Setting up a DAG hence means also setting up these variables which either are well-defined in some documentation or need to be communicated. It would be easier to have a protocol in place as part of the DAG template which defines what the variables are and what value they need.
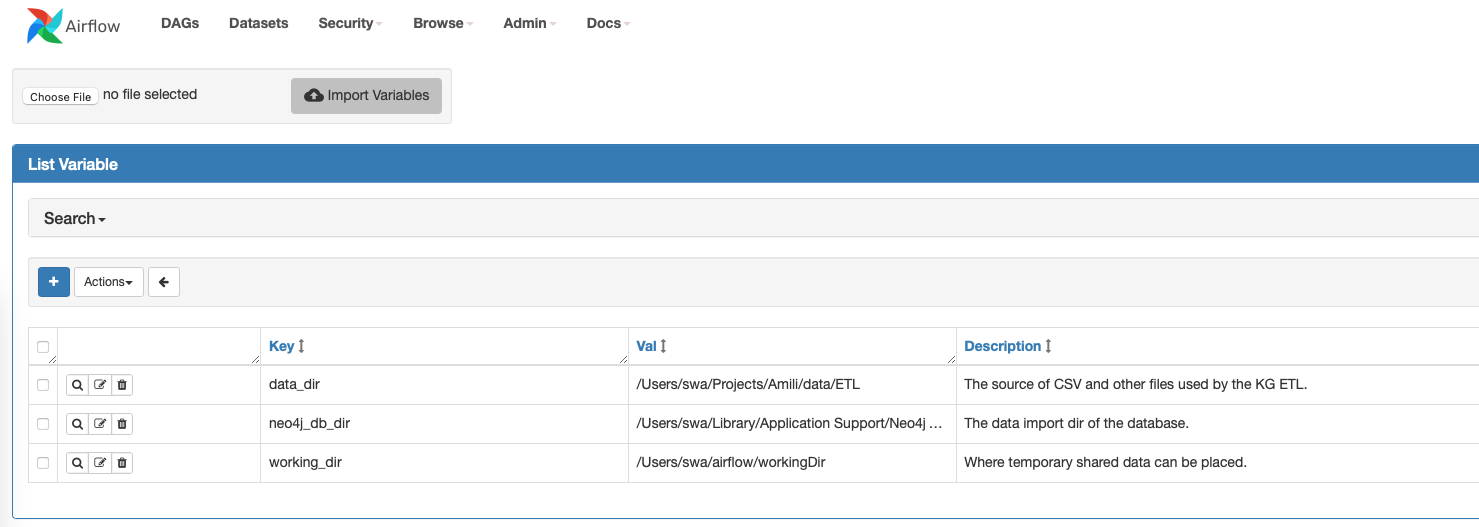
A separate dashboard of the DAG outputs would be also a great thing to have but this demands some custom web development (accessing the Airflow Rest API).
Local setup
There are various ways you can set up Apache Airflow locally:
- an easy way is via the Astronomer CLI but it’s a custom Airflow solution which focuses on the commercial Astronomer wrapper
- this docker compose template
- a standard Conda environment setup.
The standard Conda setup is the preferred way to go because it allows easier setup of packages, access to configuration and, well, full control in general:
- create a normal conda env, something like
conda create --name airflow python=3.9 - install airflow in it
- initial a database (
airflow db init)
The lightweight database solution out of the box is Sqlite but it’s advized to set up Postgresql.
Once this is set up you have a dag-directory wherein you can dump flows. The easiest way to test them is via
airflow dags test your-dag
which runs the flow in the same way as you would trigger it via the UI but it does not leave a log trace in the database. This does allow one to debug things with breakpoints and all that. If you want to debug things that way you need to refactor your Python code and debug/unit-test things like any other Python script.
Neo4j provider
Airflow has a Neo4j provider but it’s a lightweight implementation lacking the necessary bits to create an ETL flow. The ETL developed below uses the standard Bolt Neo4j driver and it needs to be installed with
pip install neo4j
Overview of an Airflow ETL
As mentioned above, loading several Gb of CSV data into Neo4j via the standard Cypher (either Create or Load CSV) path takes days. This approach works well for incremental changes and near realtime changes. The only fast and efficient way to load a large amount of data is via the neo4j-admin utility. It means, however, that
- it erases any existing data
- you need access/permission to the executable
- you need to restart the Neo4j database (and often explicitly restart it via Cypher as well).
So, although this batch import has been developed for Airflow, it still requires a manual post-fix.
The various operators (DAG steps) are on their own useful for similar jobs and the whole flow is demonstrative of how things should be organized in general.
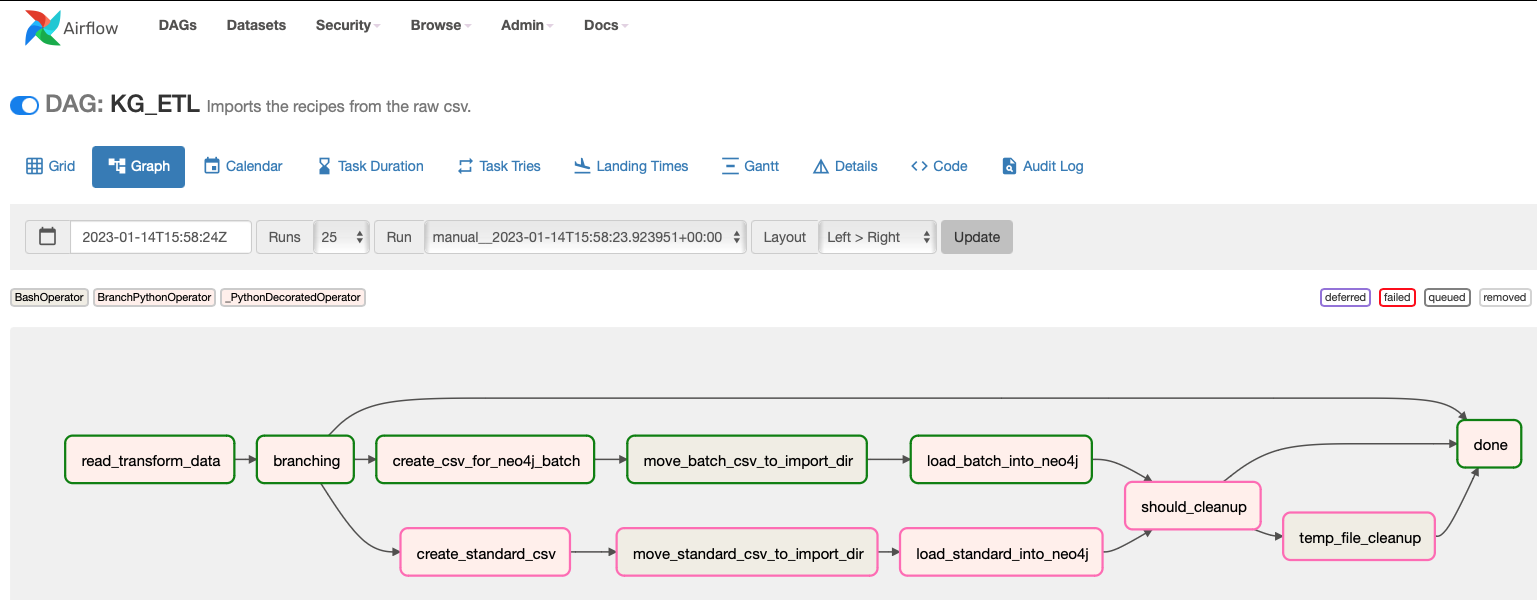
The starting point is a PyArrow Feather file. It takes just a couple of lines to convert a CSV to a Feather file and it decreases the file size tremendously. You can use Parquet or even the original CSV but it’s clear that CSV is a wasteful format.
The ETL can either load data via the aformentioned LOAD CSV way or via the neo4j-admin (batch) utility. This decision is made on the basis of the configuration.
# ============================================================
# ETL of sone raw data.
# - topn: take only the specified amount of the data source (default: -1)
# - transform: whether to transform the raw data or use the (supposedly present) clean feather file
# - import: can be 'none', 'batch' or 'load' (default: 'load'). The batch means that the batch CSV format is created for use with the neo4j-admin import CLI. The load means the Cypher LOAD CSV will be used.
#
# The following variables are expected:
#
# - data_dir: where the source data is
# - working_dir: a temporary directory
# - neo4j_db_dir: the root directory of the Neo4j database
# ============================================================
@dag(
schedule="@once",
start_date=datetime(2023, 1, 1),
catchup=False,
default_args={
"retries": 1
},
dag_id="KG_ETL",
description="Imports the the raw csv.",
params={
"topn": -1,
"transform": True,
"import": "batch",
"cleanup": False
},
tags=['Xpertise'])
The default configuration can be overriden when testing like so
airflow dags test kg_etl --config="{'import':'load'}"
The configuration helps to run part of the flow. For example, if you want to only transform part of the raw data to the necessary CSV files (nodes and edges) you can use
airflow dags test kg_etl --config="{'topn':1000, 'transform':true, 'import':'none'}"
This takes the first 1000 rows and creates nodes.csv and edges.csv without importing anything in Neo4j.
The reading and transformation phases are standard Pandas operations and data wrangling. Neo4j needs in all cases the three CSV files for import and they have to sit in the import directory of the database. So, this database directory is necessarily a parameter of the DAG. The extra step necessary is to copy/move the generated files to this directory. This bash operation is either a simple cp or ssh (scp) command depending the topology of the solution.
Once the files are in the database directory they can be loaded in one of the two ways (batch or load). This is where either a Bolt connection is set up or where the neo4j-admin utility is called.
Directories
The following directories have to be configured on Airflow
- datadir: the source of data (CSV or Feather file)
- workingdir: a temporary directory
- neo4jdbdir: the root of the database. This directory contains underneath the
bin/neo4j-adminand theimportdirectory. Neo4j will not import from anywhere else but this directory, unfortunately. It is possible to usehttp://rather thanfile://but with very large files this is not practical.
Directory structure
The organization can be used as a template for all Airflow efforts:
- operators Contains the DAG operators
- shared Contains the shared Python functions, constants and alike
- main.py The main DAG
- requirements.txt The Python package dependencies in a classic pip format
- variables.json The variables which have to be defined in Airflow
- connections.json The connections which have to be defined in Airflow.
The requirements, variables and connections should be used by a CI/CD pipeline to set things during deployment to Airflow.
Main DAG
The way one defines a flow (see the diagram above) in Airflow is somewhat idiosyncratic. The main file contains the necessary preambles as well as the flow definition:
@dag(
schedule="@once",
start_date=datetime(2023, 1, 1),
catchup=False,
default_args={
"retries": 1
},
dag_id="KG_ETL",
description="Imports the graph from the raw csv.",
params={
"topn": -1,
"transform": True,
"import": "batch",
"cleanup": False
},
tags=['Xpertise'])
def flow():
etl = read_transform_data()
what = which_import
load_standard = load_standard_into_neo4j()
load_batch = load_batch_into_neo4j()
move_standard_csv = move_standard_csv_to_import_dir
move_batch_csv = move_batch_csv_to_import_dir
standard_csv = create_standard_csv()
batch_csv = create_csv_for_neo4j_batch()
clean_up_decision = should_cleanup
end = done()
etl >> what
what >> standard_csv >> move_standard_csv >> load_standard
what >> batch_csv >> move_batch_csv >> load_batch
load_standard >> clean_up_decision
load_batch >> clean_up_decision
clean_up_decision >> temp_file_cleanup
temp_file_cleanup >> end
clean_up_decision >> end
what >> end
flow = flow()
The names used in this flow definition are one-to-one with the operators defined. These operators are in essence just Python function and bash commands but do consult the docs for the many operator you can engage in a flow.
Variables
The variables.json defines the variables which have to be set in Airflow. The format is straightforward and needs to be used by CI/CD during deployment.
{
"data_dir": {
"value": "/Users/me/Projects/ETL",
"description": "The source of CSV and other files used by the KG ETL."
},
"neo4j_db_dir": {
"value": "/Users/me/Library/Application Support/Neo4j Desktop/Application/relate-data/dbmss/dbms-b8ef492f-0c84-4b56-8d83-a6d4f3b800e0",
"description": "The data import dir of the database."
},
"working_dir": {
"value": "/Users/me/temp",
"description": "Where temporary shared data can be placed.",
}
}
These variables are accessed in a flow like this:
working_dir = Variable.get_variable_from_secrets("working_dir")
Connections
Just like variables, a connection is a setting defined in Airflow which can be accessed inside the operators.
The ETL uses only the knowledge-graph connection to Neo4j:
{
"knowledge-graph": {
"id": "knowledge-graph",
"host": "super-secret.neo4j.io",
"schema": "neo4j+s",
"login": "neo4j",
"password": "neo4j",
"port": 7687,
"type": "neo4j"
}
}
and it can be accessed like so in the DAG:
def get_connection():
con = Connection.get_connection_from_secrets("knowledge-graph")
print( f"{con.schema}://{con.host}:{con.port}")
Requirements
The requirements files is like any Python project a set of packages:
neo4j
pandas
numpy
...
It should be used in a CI/CD pipeline to set up the Python environment.
This automatically brings up the issue of clashing packages for different flows. The way this can be resolved is via the @task.virtualenv attribute, for example
@task.virtualenv(
task_id="virtualenv_python", requirements=["colorama==0.4.0"], system_site_packages=False
)
def callable_virtualenv():
"""
Example function that will be performed in a virtual environment.
Importing at the module level ensures that it will not attempt to import the
library before it is installed.
"""
from time import sleep
from colorama import Back, Fore, Style
print(Fore.RED + "some red text")
print(Back.GREEN + "and with a green background")
print(Style.DIM + "and in dim text")
print(Style.RESET_ALL)
for _ in range(4):
print(Style.DIM + "Please wait...", flush=True)
sleep(1)
print("Finished")
virtualenv_task = callable_virtualenv()
In addition, one can also use the following operators to have a clean separation:
- PythonVirtualeEnvOperator – this one will build new virtualenv every time it needs one so might be a little brittle
- KubernetesPodOperator – where you can have different variant of the images with different environments and choose the one you want for each task (requires Kubernetes)
- DockerOperator – same as KubernetesPodOperator, but requires just Docker engine
Of course, there is also the option to access lambda function and whatnot in the cloud.
ETL Operators
There are some complex things in Airflow but this example is to show that things can be also quite easy. The DAG step to move files from one place to another looks like this:
# this is the source data
data_dir = Variable.get_variable_from_secrets("data_dir")
# this is the temp data
working_dir = Variable.get_variable_from_secrets("working_dir")
# the dir of the Neo4j database
neo4j_db_dir = Variable.get_variable_from_secrets("neo4j_db_dir")
neo4j_import_dir = os.path.join(neo4j_db_dir, "import")
move_standard_csv_to_import_dir = BashOperator(
task_id="move_standard_csv_to_import_dir",
bash_command= f"""
mv '{os.path.join(working_dir,node.csv)}' '{neo4j_import_dir}' && mv '{os.path.join(working_dir, edges.csv)}' '{neo4j_import_dir}'
"""
)
Batch loading of the CSV files is also quite simple
@task()
def load_batch_into_neo4j(**ctx):
cmd = BashOperator(
task_id='csv_batch_load',
bash_command='bin/neo4j-admin database import full --delimiter="," --multiline-fields=true --overwrite-destination --nodes=import/nodes.csv --relationships=import/edges.csv neo4j',
cwd=neo4j_db_dir
)
cmd.execute(dict())
say("Batch load done. Please restart the db in order to have the db digest the import.")
# possibly requires a 'start database neo4j' as well
The tricky part resides in the correct parametrization and orchestration of tasks like these.
The actual data wrangling is really unrelated to Airflow is like any other Pandas effort. You can do all the hard work in Jupyter and paste the result in a task, for example:
@task()
def create_standard_csv(**ctx):
"""
Creates the CSV files for LOAD CSV via cypher.
"""
ti = ctx["ti"]
params = ctx["params"]
# ============================================================
# Load data
# ============================================================
clean_feather_file = ti.xcom_pull(
key='clean_feather_file', task_ids='read_transform_data')
if clean_feather_file is None:
raise Exception("Failed to get the clean_feather_file path.")
if not os.path.exists(clean_feather_file):
raise Exception(f"Specified file '{clean_feather_file}' does not exist.")
df = pd.read_feather(clean_feather_file)
logging.info("Found and loaded clean data")
# ============================================================
# Nodes
# ============================================================
nodes_file = create_nodes_csv(df, False, True)
ti.xcom_push("nodes_csv_file", nodes_file)
logging.info(f"Nodes CSV saved to '{nodes_file}'")
The create_nodes_csv call is where you can paste your Jupyter wrangling code. The XCOM push and pull methods is Airflow’s way to exchange (small amounts of) data between tasks. Here again, I think that the terminology is awkward, it obfuscates adoption and understanding.
Closing thoughts
All of the Apache frameworks and tools have the same mixture of good-bad (or love-hate if you prefer) and it always takes some time and energy to learn the ins and outs. Airflow is a stable ETL platform and if Python is your programming language it’s a great open source solution. Like any OSS it requires learning and additional embedding efforts.
Personally, I very much enjoy working with Airflow and would recommend it to any customer in need of an ETL solution and a Neo4j CDC or data ingestion need in particular.


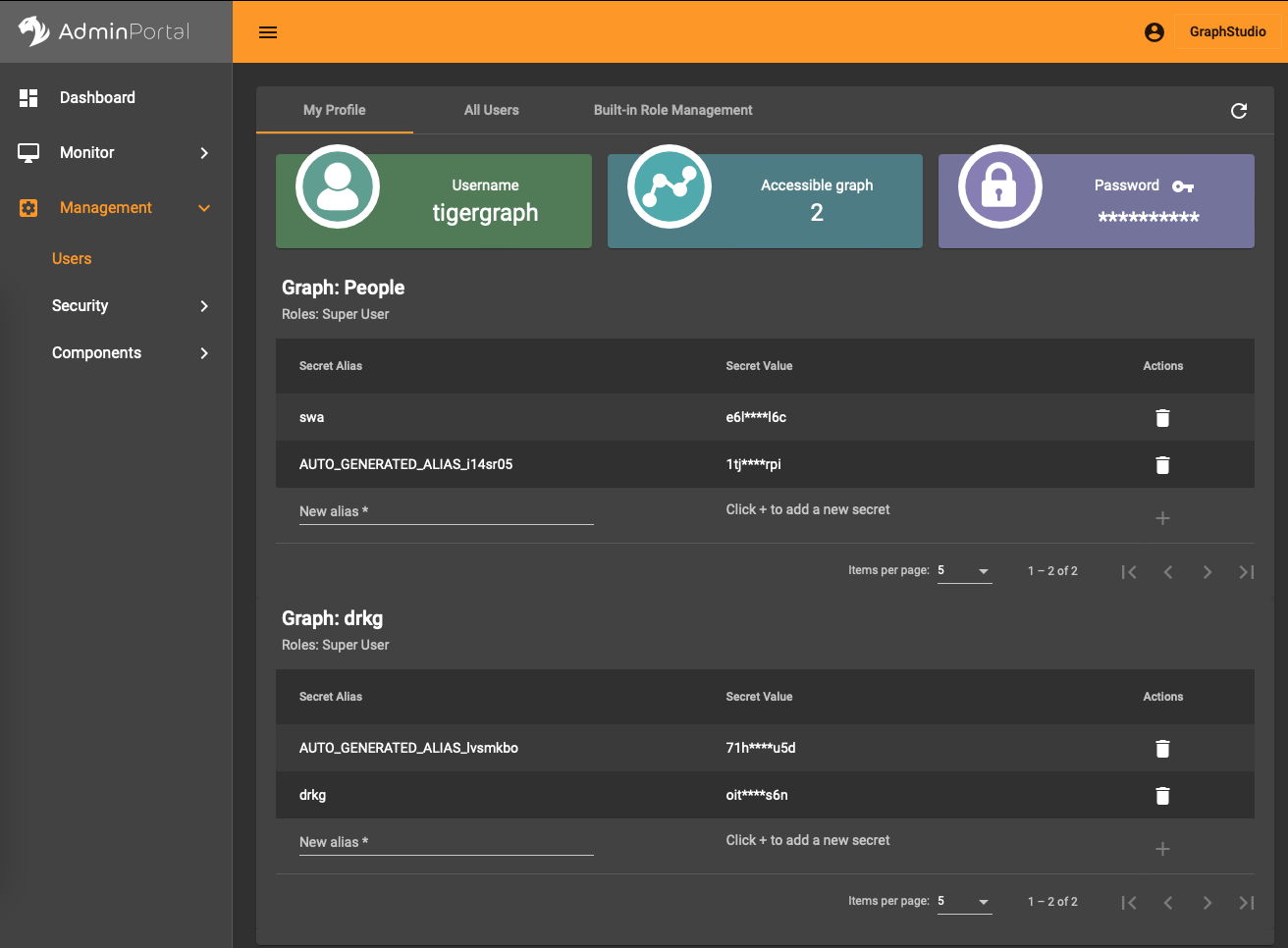 In the AdminPortal (see image) you should add a secret specific to the database. That is, you can’t use a global secret to connect, you need one per database.
In the AdminPortal (see image) you should add a secret specific to the database. That is, you can’t use a global secret to connect, you need one per database.